Our lab uses a diverse suite of approaches to uncover the processes shaping species formation and patterns of genomic diversity. We work on three main key areas/systems.
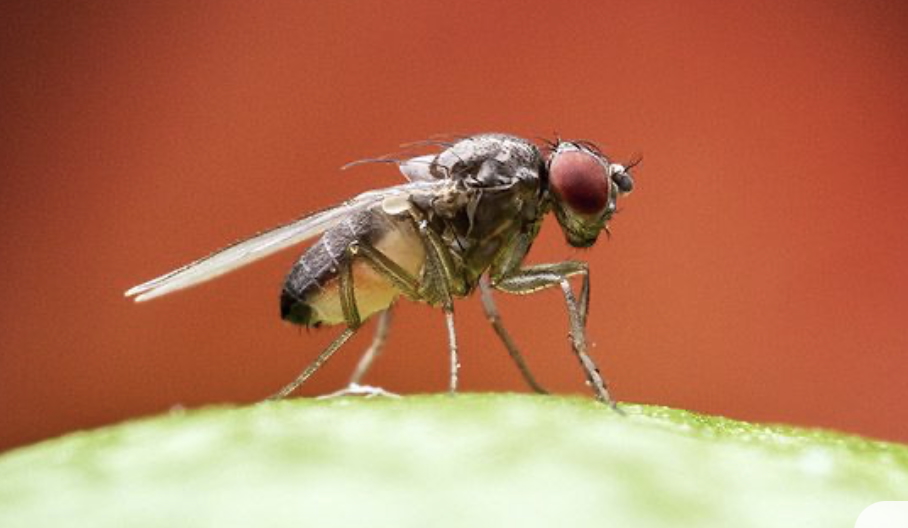
Uncovering Genomic Mechanisms Driving Species Divergence in Drosophila
We study the roles of gene flow and natural selection during speciation, and the role of inversions on species divergence. Our work is focused on the divergence between Drosophila pseudoobscura and its close relatives D. persimilis and D. p. bogotana. The presence of chromosomal inversions in this species group precludes fine scale genetic analyses to characterize the genetic basis of species isolation mechanisms or phenotypic differences between species. In collaboration with Dr. David Stern (HHMI), we have developed genetic tools and reagents specific for this species group that are being used in genome editing experiments aimed at generating/reverting chromosomal inversions. In addition, we are studying the evolution and functional genomics of conserved long non-coding RNAs (lncRNAs) and small ORFs in Drosophila. These understudied genomic elements appear to have important roles in organismal biology and particularly in the divergence of male reproductive processes.
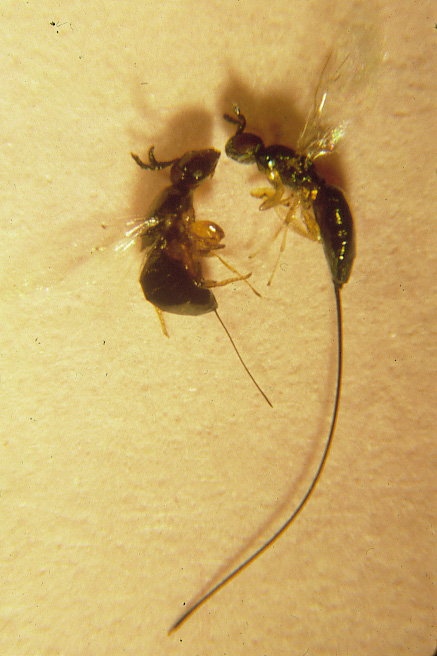
Investigating Plant-Insect Co-Evolution Through the Fig and Fig Wasp Mutualism
Both ecologically and evolutionarily, mutualisms represent one of the most influential of all biological interactions, with fundamental consequences for the evolution and maintenance of biodiversity. Figs (Ficus sp., Moraceae) and their pollinating wasps (Agaonidae, Chalcidoidea, Hymenoptera) constitute one of the most extraordinary pollination mutualisms known. In addition to the pollinating wasps, a diverse community of non-pollinating wasps also exploits figs. We are using a diverse suite of integrative “omic” and ecological approaches to characterize the geographic context co-diversification of this mutualism, to understand patterns of genome diversity and population genetic diversity in fig wasps, and to uncover genetic and morphological innovation involved in the evolution of synergistic and antagonistic interactions in this mutualism.
Studying the Genomic Basis of Adaptation in Human Parasites, and Plant Domestication
Although these topics are not our current research focus, we are happy to host students interested in plant domestication (mostly figs now, and rice in the past) or human parasite evolution (mostly Trypanosoma cruzi, the agent of Chagas’ disease). We have studied the evolutionary history of T. cruzi and other Trypanosomatids using population genetic and genomic data, uncovering evidence of hybridization as well as a new genetic lineage endemic to North America. Using comparative genomic data, we have shown that inferred genome-wide signals of positive selection are higher in T. cruzi proteins than in Leishmania spp. proteins, a result consistent with the greater versatility of T. cruzi in its host range, cell tropism and cell invasion mechanisms. Currently, we are using genomic data to expand our understanding of T. cruzi diversification, and to the study the dynamics of gene duplication in Trypanosomatid genomes.